|
|
Human induced pluripotent stem cell (iPSC)-based models are a valuable resource for studying disease mechanisms in vitro at the cellular level[1], screening potential new therapeutics[2], and investigating the propensity and mechanism for the development of toxic side effects caused by a drug treatment[3]. Such iPSC-based models enable research to be performed under defined experimental conditions and in a reproducible manner.
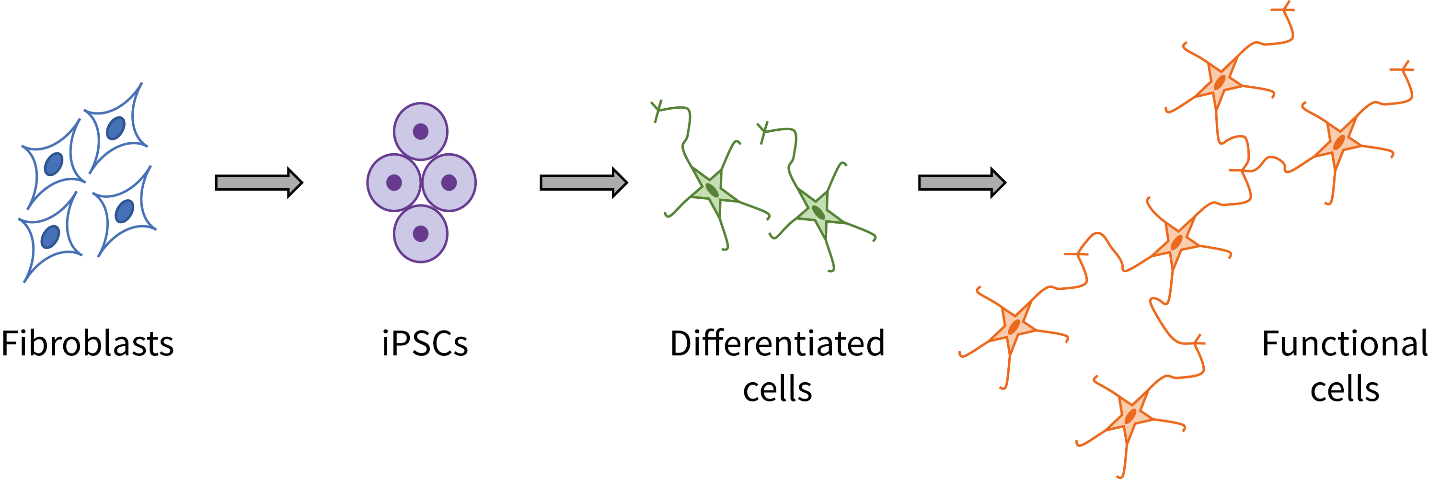
Creation of iPSC-based disease models typically involves multiple steps, each optimized for a particular disease and cell type. From obtaining primary target cells, through generation of iPSCs, differentiation of iPSCs into disease cell types, culturing differentiated cells to develop the disease phenotype in vitro, and finally, treating the cell model with test agents and evaluating changes in phenotype, the process is time consuming and requires special skills.
How can I access diseased tissues and generate an iPSC-based disease model?
One of the main challenges in developing iPSC-based disease models is the first step: access to human tissue with the disease phenotype of interest. Whether studying cellular mechanisms of certain diseases or screening for potential new drugs for therapeutic benefit, the cells used in in vitro models should reflect the genotype and phenotype of the diseased cells in vivo.
Two general approaches can be used to overcome this challenge.
- Procurement of tissue from individuals with a particular disease phenotype
- Gene-editing approaches – such as CRISPR-Cas9 – where cells from a “healthy” donor are converted to the disease genotype of interest.
Procurement of tissue
The most straightforward way to obtain relevant cells for a disease model is to target patients with the disease of interest and obtain a source material, such as blood, urine or skin, from which to isolate primary cells to reprogram (Figure 1). This approach is advantageous in that all of the genetic changes associated with a disease phenotype are incorporated within the primary cells and will therefore be included in the derived disease model. However, disease-affected tissue can often be difficult to obtain, especially for rarer conditions and disorders. Additionally, due to the diseased nature of certain tissues, deriving a viable primary cell culture for reprogramming can be problematic.
Gene-editing
Introducing disease-causing genetic abnormalities into healthy iPSC, through gene editing technologies such as CRISPR-Cas9, is an alternative strategy (Figure 2). This has the advantage that specific genetic changes can be introduced and manipulated to assess disease mechanisms and treatment responses, and, through the use of healthy donor tissue, negates the need to obtain tissue from disease affected individuals. Such genetic manipulation to create a disease model requires expert knowledge and is often a protracted process, but it does allow good control of the overall genetic background for comparison with the wild type control.

Figure 1. CRISPR-Cas9 gene editing complex. The CRISPR-Cas9 complex includes the Cas9 Nuclease (“Scissors”) and a targeting sgRNA that directs the complex to the appropriate place on the genome.
How can REPROCELL help you generate a patient-derived, iPSC-based disease model?
Through our extensive Global Tissue Network and alliances, REPROCELL can source a variety of diseased and healthy tissue samples (for example skin, blood or urine cells) for use with REPROCELL reprogramming technologies to generate iPSC lines on behalf of our clients. Tissues and cells that we source are fully anonymized and consented for use in all types of iPSC-based projects, and they come with valuable donor demographic information. We can also perform whole genome sequencing or molecular characterization of tissue or primary cells to provide further genetic information or confirm disease-relevant mutations.
Table 1 shows a few of the disease areas available through our Tissue Network. We can obtain the tissues, isolate primary target cells, reprogram using our state-of-the-art StemRNA 3rd Gen Reprogramming Technology, and deliver donor-derived iPSCs ready for use. As our network is constantly expanding, please check with us to find out if we can source your target cell type or tissue.
Table 1: Examples of Tissue Network disease area

How can REPROCELL help you generate a gene-edited, iPSC-based disease model?
CRISPR-based gene editing technology is revolutionizing the biopharma industry, enabling advances in disease modeling that were impossible just a few years ago. The use of techniques such as CRISPR enable precise, directed creation of knock-outs and knock-ins (including single base changes) in many cell types. REPROCELL provides CRISPR-Cas9-based gene editing services.
CRISPR technology generally involves a 3-step process. First, the design of guide RNAs and choice of optimal nuclease. Second, the delivery of components to the cell to allow editing to occur. Third, screening and analysis of the generated clones and their frequency to enable selection and expansion of the desired edit. The third step is the most time-consuming and labor-intensive, taking many weeks to complete when using the standard method of limiting dilution, colony picking, and sequence screening. In contrast, REPROCELL’s CRISPR-SNIPER technology uses the proprietary SNIPER method to efficiently and accurately screen clones, thereby significantly streamlining and improving the selection process.
What is SNIPER?
SNIPER (Specification of Newly Integrated Position and Exclusion of Random-integration) uses a combination of long, high-stringency donor DNA with a digital PCR-based quantification method and empirical culture condition determination to prescreen and identify colonies most likely to be positive for the desired change. As a result, fewer clones need to be selected for sequence verification, and the success rate for isolating homoallelic and heteroallelic clones with the correct genetic change is greatly increased compared to conventional methods.
SNIPER can be used with knock-in, knock-out and single nucleotide base change edit methods with a high degree of efficiency. For single nucleotide base change generation, where only the desired base change is required, use of a traditional screening gene, such as PuroR or GFP to aid selection, is not an attractive option. Consequently, for standard base change protocols, screening of large numbers of colonies is required to identify positive clones, resulting in a significant investment in time and resources. Addition of SNIPER to single base change modification screening results in a 100-fold or greater increase in the fraction of positive clones, greatly improving this resource-intensive step.
Table 2. Comparison of screening methods for a typical knock-in study

To learn more about the CRISPR-SNIPER technology, please see this recent webinar.
CRISPR-SNIPER Gene Editing Advantages
CRISPR-SNIPER gene-editing has several advantages over more traditional CRISPR-Cas9 methods making it the preferred choice for developing an iPSC-disease model for research.
- Improves efficiency of editing in iPSC lines
- Accelerated identification of positive clones
- Delivers high success rate for single base pair change modifications for disease model generation
- Great for knock-out (KO) and knock-in (KI) applications
- Provides Freedom-to-Operate for research use including drug screening
Such advantages make CRISPR-SNIPER-edited iPSC lines a valuable, relevant, and comparative alternative to iPSC lines directly derived from a diseased individual.
Case Study
Our Alzheimer’s disease case study demonstrates both approaches.
Case Study: Creation of Alzheimer’s Disease Models
To create an Alzheimer’s Disease (AD) neuronal disease model, we utilized both our global Tissue Network and gene-editing capabilities. Wild type (WT) iPSCs were generated from a healthy donor, and CRISPR-SNIPER gene editing used to introduce a mutation in the AD-associated gene, PS1. In parallel, an iPSC line bearing a mutation in the AD-associated gene, PS2, was generated from tissue donated from a donor with AD. All three iPSC lines – WT, PS1 and PS2- were treated with retinoids to generate neurospheres[4] and subsequently cultured in the presence of mitotic inhibitors to generate neurites[4]. Comparison of neurite characteristics demonstrated that neurospheres from iPSC lines with the different genetic backgrounds displayed quantifiable differences in neurite length and number.


iPSC-derived Alzheimer’s disease model Neurospheres
Neurospheres were created from iPSCs with a healthy (WT) genotype, as well as from an AD patient with a PS2 mutation, or by editing the WT to create a PS1 mutation.
Make REPROCELL your iPSC-based disease model partner
REROCELL is your preferred partner throughout the entire disease model creation process. To find out more about the REPROCELL Tissue Network, Reprogramming Services, CRISPR-SNIPER Gene Editing and Molecular Services, please contact us.
References
- Sarkar et al. Efficient generation of CA3 neurons from human pluripotent stem cells enables modeling of hippocampal connectivity in vitro. Cell Stem Cell22:684-697 (2018)
- Czerniecki, et al. High-throughput screening enhances kidney organoid differentiation from human pluripotent stem cells and enables automated multidimensional phenotyping. Cell Stem Cell 22:929-940 (2018)
- Holmgren et al. Expression profiling of human pluripotent stem cell-derived cardiomyocytes exposed to doxorubicin – Integration and visualization of multi-omics data. Tox Sci 163: 182-195 (2018)
- Clarke et al. A robust and reproducible human pluripotent stem cell derived model of neurite outgrowth in a three-dimensional culture system and its application to study neurite inhibition. Neurochem Intl 106:74-84 (2017)



















Tell Us What You Think!