|
|
Extracellular vesicles (EVs) are known for their role in facilitating wide range of intercellular communication such as regulation of the immune response and stem cells differentiation.
What is an extra-cellular vesicle?
EVs are membrane-enclosed particles ranging in size from 30nm to 300nm and their characteristic depend on the cell type releasing them. Often they will carry cargo of active molecules like RNA, DNA, proteins, metabolites and lipids. Moreover, EVs can be found in most biological fluids with the ability to cross biological barriers, and in the case of pathological conditions, the composition of the cargo will be distinctive to the disease. Altogether, these features have led to intense research into the potential role of EVs for diagnostics, biomarkers and as vehicles for drug delivery.
EV characterization
Due to the small size of EVs, often below the diffraction limit of light (~250nm), a full characterization of their size and cargo, as well as the dynamic processes of cellular uptake and release pose a major challenge.
The most commonly used techniques to study EVs are electron microscopy (EM) for visualization, and flow-based methods to estimate the size and concentration. The main disadvantage with EM is that the sample preparation can affect both the morphology and the composition of the EVs and that it is difficult to characterize the distribution and number of biomarkers. Flow-based methods come with different disadvantages mostly due to the difficulty of detecting EVs on the lower end of the size spectrum, discrimination between sub-population of EV and data interpretation can be difficult.
Recent advances in the field of fluorescence microscopy with the development of single-molecule localization methods have provided scientists with unique tools to acquire high-resolution images of EVs, estimate the size and concentration, as well as the possibility to study their behavior in biological contexts.
Super-resolution imaging of EVs with ONI’s Nanoimager
EV populations are heterogeneous both in size and composition, and these differences within a population are most likely responsible for different biological outcomes. In order to provide scientists with an easy-to-use and powerful instrument to fully investigate the characteristics of individual EVs, ONI has developed a bench-top compatible super-resolution microscope with single-molecule sensitivity, the Nanoimager.
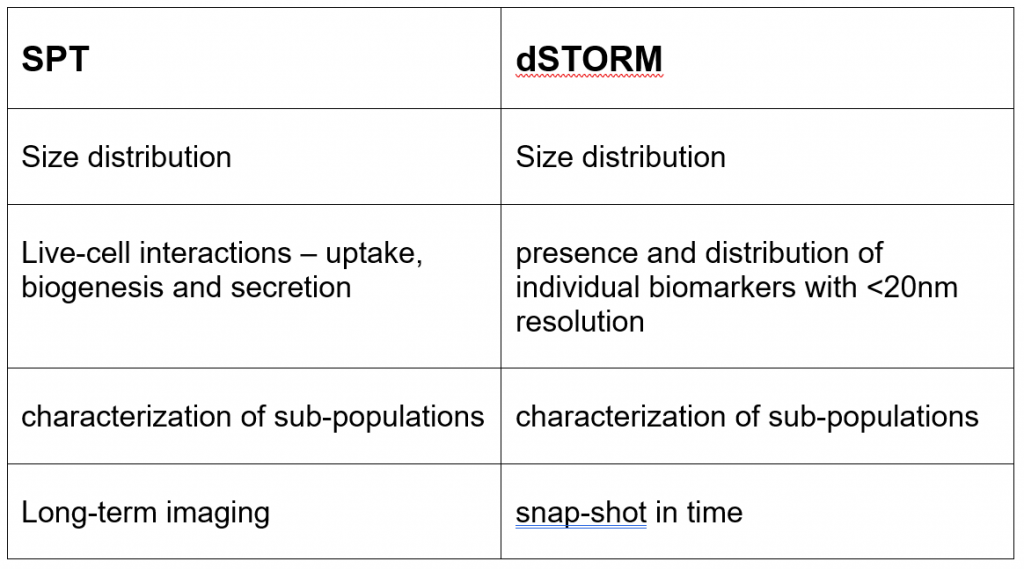
The ideal super-resolution technique to resolve individual membranes and biomarkers with a resolution of up to 20nm is dSTORM (direct STochastic Optical Reconstruction Microscopy). Figure 1 shows a side by side comparison of an EV image acquired in a wide-field mode and the reconstructed dSTORM image. The ability to measure the size distribution of EVs in a population, as well as visualizing the presence and distribution of individual biomarkers on a single-EV level, allow the detailed characterization of sub-populations which, in turn, informs on the targeting specificity.
dSTORM is a powerful method to access structural and interaction information at a biologically relevant scale. However, the resulting image is a snap-shot in time and does not provide us insights into the dynamic nature of the EV cycle. To address this, another technique, single-particle tracking (SPT) can be used to visualize and quantify EV behavior.
Characterizing EVs size, cargo and uptake with Single-Particle tracking (SPT)
SPT is a method by which individual molecules are followed over a period of time to extract dynamic information. By imaging and tracking fluorescently labeled EVs in solution, and measuring the diffusion rate you can extract information related to size. SPT can also be carried out in a dual-color mode, in which case the size of a specific EV can be correlated with the incorporation of a specific biomarker. SPT on single EVs in a cellular context provides valuable dynamic information on the interaction and uptake by cells, as well as their intracellular fate.
Figure 2 shows the uptake and subsequent tracking of single EVs in live cells. The directionally and the diffusion coefficient of the EVs provide valuable information about their intracellular behavior. Parallel labeling of intracellular components would further inform on their intracellular fate.
The Nanoimager enables researchers to combine super resolution imaging with single-particle tracking, making it one of the most specialized tools for EV characterization. Unlike other super-resolution microscopes, the Nanoimager is as big as a desktop computer and can be installed anywhere (no need for an optical table or a dark room). Thanks to a high-sensitivity camera, uniquely designed optics, live-cell compatibility and user-friendly software specifically designed for SPT and dSTORM, the Nanoimager provides a unique opportunity for all scientists to access biological information on a single-molecule level.
Taking EV characterization to the next level with ONI’s Nanoimager – Figures
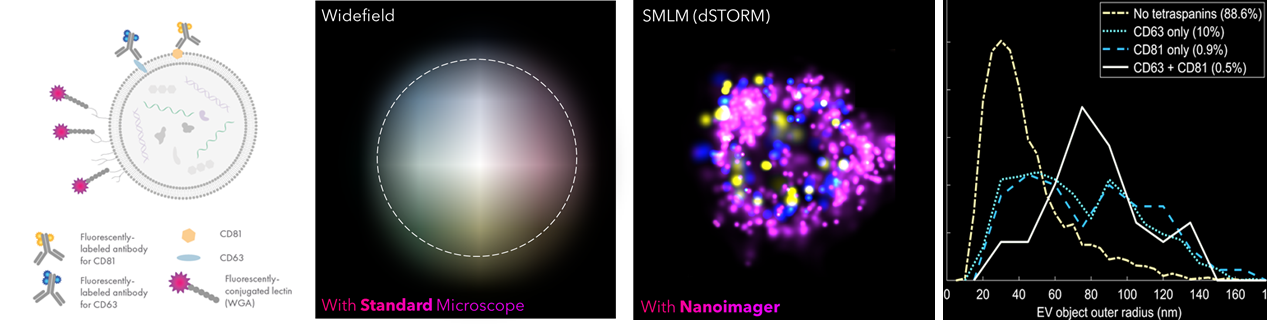
Figure 1. dSTORM imaging and characterization of EVs. Experimental design, imaging, and biomarker distribution of EVs from keratinocytes. A. Fluorescence labeling schematic B. Widefield image of a single EV carrying a WGA membrane (purple) and the tetraspanins CD63 (blue) and CD81 (yellow). C. Reconstructed dSTORM image D. Biomarker distribution as a function of the EV radius
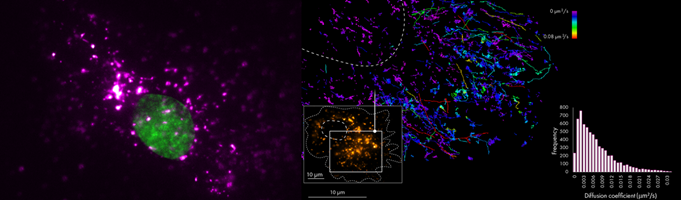
Figure 2. Live-cell imaging of EVs. A. Live-cell image of Burkitt’s lymphoma cell. The nucleus is labeled with Hoechst (green) and EVs with AF647 (purple) B. Single-particle tracking of EVs labeled with AlexaFluor488. Tracks are colored according to the diffusion coefficient.


Tell Us What You Think!