|
|
Highlights
- The biggest opportunity for success in cardiac drug discovery and development is to make these processes more efficient and affordable, since they carry relatively higher risk of clinical trial failure compared to other disciplines.
- Human iPSC technology is increasingly utilized to develop disease models that reliably mimic the diversity of patient pathogenicity and disease severity in vitro.
- Phenotypic screening based on iPSC-derived disease models is making cardiac drug discovery more time and cost effective by enabling target-agnostic studies and increasing the translational power of early drug discovery outcomes.
Background
Since 1960, there has been a huge growth in treatments for cardiac diseases 1. Although they have increased the longevity of human population, cardiac pathologies remain leading causes of morbidity and mortality worldwide 1,2. For a number of reasons, the number of cardiovascular drugs entering clinical trials has been in decline since 1990 3. The poor translation of safety and efficacy from preclinical models to humans has caused several major failures in advanced stages of clinical trials in the past, resulting in a challenging regulatory environment, compared to most other therapeutic areas. Therefore, phase III clinical trials for cardiac diseases now need to be performed with relatively larger groups of patients and need to demonstrate the efficacy in longer-term follow-up than other disciplines 4. Altogether, these factors cause a huge increase in the overall costs of cardiac drug discovery and development.
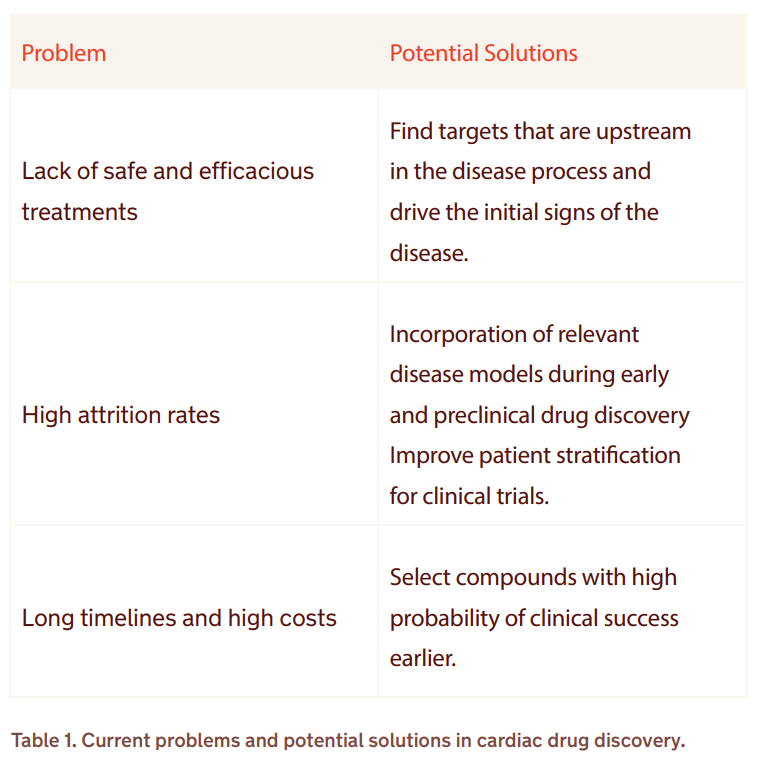 Despite having available treatments for most cardiac diseases, many of the approved drugs target the overall symptoms of the disease, instead of the underlying causes. Moreover, there are large groups of patients for whom current treatments are not sufficiently effective4. There is a need to find better and more efficacious treatments, but to do so, it is important to make the process more affordable and de-risk potential clinical candidates at early stages to optimize the use of resources (Table 1).
Despite having available treatments for most cardiac diseases, many of the approved drugs target the overall symptoms of the disease, instead of the underlying causes. Moreover, there are large groups of patients for whom current treatments are not sufficiently effective4. There is a need to find better and more efficacious treatments, but to do so, it is important to make the process more affordable and de-risk potential clinical candidates at early stages to optimize the use of resources (Table 1).
The discovery of human induced pluripotent stem cells (iPSCs) by Shinya Yamanaka and Kazutoshi Takahashi in 2007, brought a new tool with a high potential to revolutionize drug discovery 5,6. iPSCs are obtained from human somatic cells, have unlimited proliferation capacity, and can differentiate into any cell type of the body, including cardiac muscle cells. Since 2010, multiple cardiac diseases have been successfully modeled using this technology 7 and it has been demonstrated that human iPSC-derived cardiomyocytes are a valuable solution for cardiac drug discovery.
In this whitepaper, we discuss the main challenges and goals in cardiac drug discovery and how human iPSCs can help achieve them.
Increasing the translational power of disease models
Preclinical testing is currently largely based on animal models, predominantly mice. Although these models have contributed towards unraveling many disease mechanisms, they are costly and present important interspecies differences to humans that diminish their predictive power for compound safety and efficacy. Murine heart rate is eight times faster than human and there are considerable differences in the expression of important sarcomere proteins, ion channel function, metabolism, and calcium handling, among others 8. For example, many mutations in the MYH7 are associated with the development of hypertrophic cardiomyopathy (HCM). MYH7 is the dominant isoform in human heart, but this is not the case for rodent models 9.
 These types of inter-species differences can lead to misinterpretations during preclinical screenings that can translate into failures during clinical trials. Human iPSC-cardiomyocytes preserve the exact same intracellular environment and genetic background of patients 10, which enables recapitulation of specific disease-linked phenotypes and underlying mechanisms (Fig.1). Human iPSC-cardiomyocytes generated from a family carrying a mutation on the gene MYH7 were shown to recapitulate the HCM phenotype in vitro, and revealed that elevation in intracellular calcium levels preceded the appearance of other phenotypic abnormalities 11. This suggested that calcium cycling alterations may be driving the disease and opened new possibilities for developing more efficacious treatments.
These types of inter-species differences can lead to misinterpretations during preclinical screenings that can translate into failures during clinical trials. Human iPSC-cardiomyocytes preserve the exact same intracellular environment and genetic background of patients 10, which enables recapitulation of specific disease-linked phenotypes and underlying mechanisms (Fig.1). Human iPSC-cardiomyocytes generated from a family carrying a mutation on the gene MYH7 were shown to recapitulate the HCM phenotype in vitro, and revealed that elevation in intracellular calcium levels preceded the appearance of other phenotypic abnormalities 11. This suggested that calcium cycling alterations may be driving the disease and opened new possibilities for developing more efficacious treatments.
In vitro models based on primary human cardiac cells could be another solution, as they are physiologically relevant. However, they are restricted by the tissue inaccessibility and lack of cell survival in culture. In addition, their capacity to discriminate between causes and effects of the disease can be limited. For example, a common consequence of atrial fibrillation (AF) is tissue remodeling. Using patients’ biopsies as in vitro models, where the remodeling is usually present, does not allow to determine the etiology of AF 12. Murine models have been generated, but they are quite resistant to developing AF under physiological conditions 13. However, human iPSC-derived cardiomyocytes can help in understanding the initial causes of AF. In a study using human iPSC-cardiomyocytes derived from patients with AF, it was shown that increased pacemaker and L-type calcium currents could be triggering specific types of AF 12.
Human iPSC-derived cardiomyocytes preserve patient’s genetic background and recapitulate specific disease-linked phenotypes, which increases their translational power.
Overall, developing more predictive in vitro disease models with human iPSCs can help towards understanding underlying disease mechanisms and identifying better targets. Therefore, decreasing the risk of failure in later stages.
Efficient screening of cardiac drug candidates
In the past, a number of challenges have hindered the incorporation of iPSCs into drug discovery projects. Differentiation procedures for human iPSC-cardiomyocytes used to be inefficient and difficult to scale up, which limited the number of cells available. Also, assays were not yet automated and miniaturized, which was needed to minimize variability and enable large screening campaigns. Nowadays, with the adequate capabilities and expertise, it is possible to obtain billions of functional cardiomyocytes from patient-derived iPSCs, thus overcoming the limitations of tissue inaccessibility and culture maintenance of primary cardiac cells.
High-throughput screening
The incorporation of human iPSC-based models has brought a new approach to drug discovery: high-throughput phenotypic screening. While traditional screenings are focused on a specific protein (target-based drug discovery), phenotypic screenings can be target-agnostic and focus on the modulation of a disease-linked phenotype 14. These screenings are more physiologically relevant and offer the possibility of identifying completely new targets or unknown mechanisms of action of known targets. For instance, human iPSC-cardiomyocytes derived from patients with a mutation in LMNA were used to model dilated cardiomyopathy (DCM) in vitro 15. The model displayed aberrant calcium homeostasis and arrhythmias, as described in the patients. Based on these DCM models, it was shown that activation of the platelet-derived growth factor (PDGF) was driving the alterations and that its inhibition led to an ameliorated phenotype in vitro 15. This described an underlying cause of the disease and identified a new potential therapeutic target for DCM.
Phenotypic screening with human iPSC-derived models can increase confidence of potential clinical candidates at early stages of drug discovery.
Clinically relevant outcomes
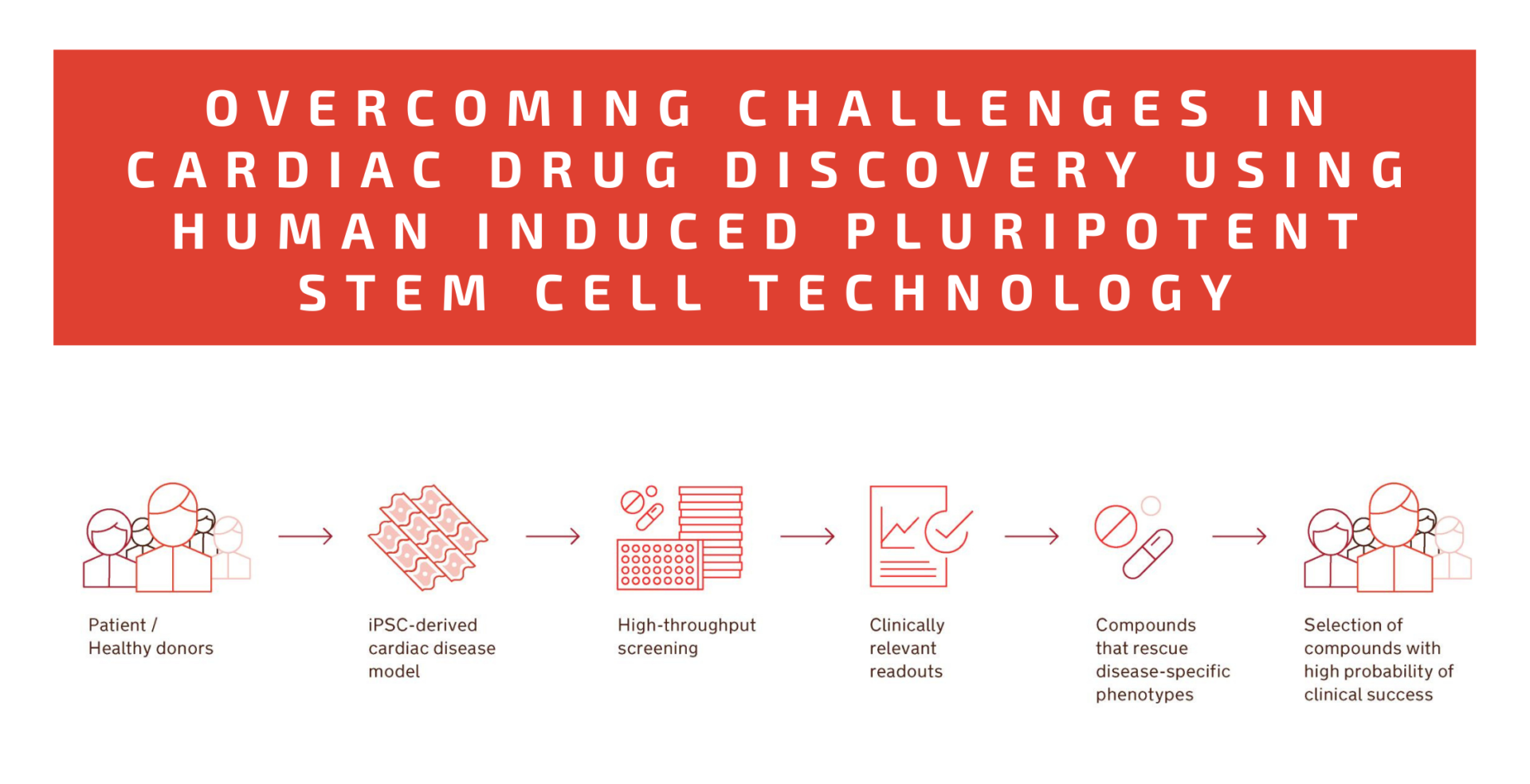
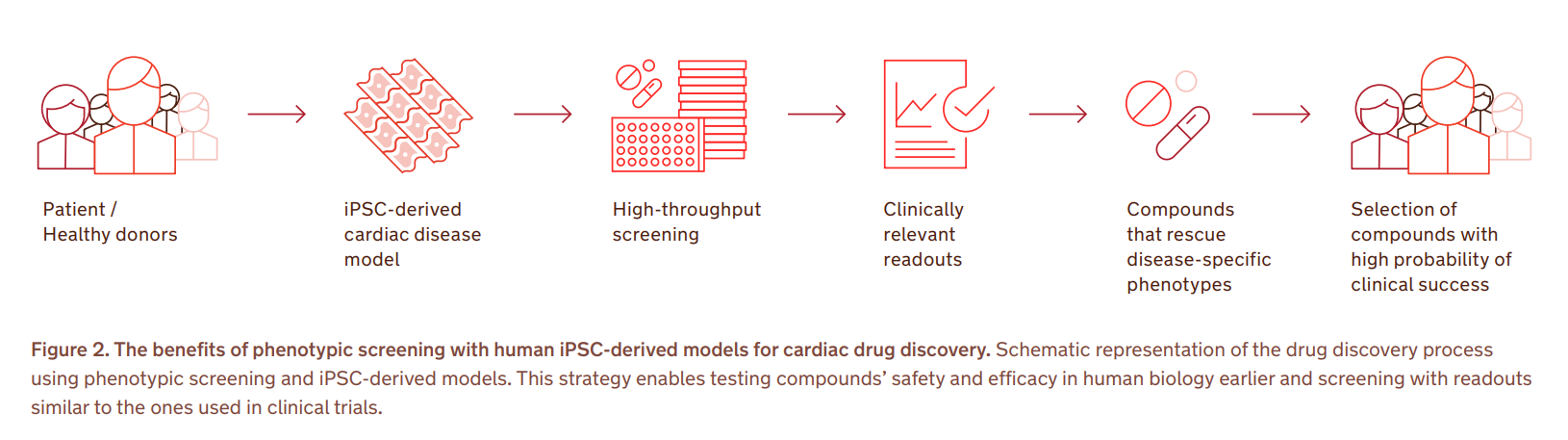
Well-designed phenotypic assays with human iPSC-based cardiac models can evaluate clinically relevant cellular functions: electrophysiology and contractile properties, metabolism, calcium handling, viability, or biomarker expression 16. This means that early drug discovery and preclinical testing outcomes are based on the same or similar parameters that are later assessed in patients, reducing the risk of failure in clinical trials and making the process more efficient (Fig. 2). Many promising lead compounds have been identified with human iPSC-based phenotypic screenings 16.
To give an example, isogenic diseased and genetically engineered human iPSC-derived cardiomyocytes were used to model long-QT syndrome (LQTS) in vitro 17. In this study, the small molecule LUF7346 was demonstrated to be a new allosteric modulator of the potassium channel hERG, with the capacity to rescue LQTS-phenotype in a human in vitro model.
More recently, human iPSC-derived models of diabetic cardiomyopathy were used as screening platform for small molecules with a phenotype-rescue approach 18. Diabetic cardiomyopathy has a complex multifactorial etiology difficult to model with traditional methods. Two iPSC models were generated, one chemically induced and one patient-derived, both recapitulated the structural and functional disarray observed in diabetic patients’ hearts. High-content phenotypic screening of 480 small molecules on the chemically induced model led to the identification of 47 hit compounds with phenotype-rescue effects. A second screening confirmed the efficacy and dose-dependency of responses for 28 hits. In a final screening with patient-derived models, two compounds were selected as the most-effective in preventing the development of diabetic cardiomyopathy phenotypes, demonstrating the power of human iPSC technology for the development of new therapeutic modalities with higher specificity and efficacy.
Effective screening of cardiotoxicity
In addition to the poor translation of efficacy from preclinical models to humans, cardiotoxicity is another major cause of drug attrition in clinical trials. In this field, human iPSC technology has proven to be a solution to predict drug-induced structural damage and arrhythmias with high efficiency. The comprehensive in vitro proarrhythmia assay (CiPA) represents a paradigm to improve the assessment of proarrhythmic risk based on mechanistic in vitro assays and in silico reconstructions of electrophysiological activity, with verification of predicted responses in human iPSC-derived cardiomyocytes. To illustrate the power of CiPA, a screening with 28 drugs with known clinical arrhythmic effects demonstrated 87% predictivity of proarrhythmic risk that was not detected in pre-clinical studies using traditional models 19.
Increased safety and efficacy for cardiac treatments can be achieve by incorporating human iPSC technology into drug discovery and development.
The future of human iPSC technology
3D models
It is important to mention that human iPSC technology is a relatively young field and is therefore still under continuous development to improve its resemblance to the complexity of the human body. One developing solution is the use of 3D engineered microtissues, which are composed of multiple cell types found within the human heart 20. These 3D models facilitate the study of cell-cell interactions and represent more complex microenvironments with higher similarity to native tissue, which may increase predictions of drug safety and efficacy even further.
For instance, 3D cardiac models better resemble the metabolism and proliferative capacity of adult hearts, which can facilitate the identification of pro-proliferative compounds for cardiac regeneration 21. In a screening with 105 small molecules and human iPSC-derived 3D cardiac organoids, novel pro-proliferative compounds with minimal side effects were identified 21. At this moment, the throughput of these models is somewhat limited compared to 2D monocultures. However, promising advancements are being made to solve this issue that will unlock broad implementation of human iPSC-derived 3D models in drug discovery 22.
Personalized and precision medicine
Some other major challenges for cardiac drug discovery are the oversimplified systems for patient stratification and the differences in mutation penetrance among patients. Patient stratification is usually based on phenotypes with different underlying etiologies, which leads to a wide range of treatment responses among patients 4,23 Studies with human iPSC technology are constantly increasing our understanding of cardiac disease etiology, setting the foundations for a mechanism-driven patient classification in the future. This will also facilitate the definition of homogenous groups of patients for clinical trials and the interpretation of clinical outcomes. On the other hand, patient-derived or genetically engineered iPSC models can help to determine the pathogenicity of specific genetic variants and predict the in vitro response to pharmacological treatments of individual patients 24,25This is of great advantage for the growing demand for personalized and precision medicine tools.
Conclusions
Cardiac drug discovery has been facing a stage of paucity, but the need for more efficacious treatments for devastating cardiac diseases is waking up the interest in this area again. The current goals in the field are challenging: finding better targets and reducing attrition rates to make the process more affordable.
Incorporating human iPSC technology in cardiac drug discovery can solve many of the current challenges in the field. This technology facilitates target identification and phenotypic screening with clinically relevant readouts. Consequently, the predictability of early and preclinical drug discovery outcomes can be increased, and therefore, the chances of success in clinical trials.
The implementation of iPSC-based models into drug discovery requires significant expertise and capabilities. To overcome these impediments and obtain high-quality results in the shortest possible timeframe, partnering with expert contract research organizations is a common option for pharmaceutical companies. Collaboration with experts avoids common pitfalls, enables selection of risk mitigation strategies, enhances productivity and, ultimately, reduces timelines and costs.
References
1. Gromo G, Mann J, Fitzgerald JD. Cardiovascular Drug Discovery: A Perspective from a Research-Based Pharmaceutical Company. Cold Spring Harb Perspect Med. 2014;4(6). doi:10.1101/CSHPERSPECT.A014092
2. Fordyce CB, Roe MT, Ahmad T, et al. Cardiovascular Drug Development: Is it Dead or Just Hibernating? J Am Coll Cardiol. 2015;65(15):1567-1582. doi:10.1016/J.JACC.2015.03.016
3. Ohlstein EH. The grand challenges in cardiovascular drug discovery and development. Front Pharmacol. 2011;JUL:125. doi:10.3389/FPHAR.2010.00125/BIBTEX
4. Figtree GA, Broadfoot K, Casadei B, et al. A Call to Action for New Global Approaches to Cardiovascular Disease Drug Solutions. Circulation. 2021;144:159-169. doi:10.1161/CIR.0000000000000981/FORMAT/EPUB
5. Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126(4):663-676. doi:10.1016/J.CELL.2006.07.024
6. Takahashi K, Tanabe K, Ohnuki M, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131(5):861-872. doi:10.1016/J.CELL.2007.11.019
7. Karakikes I, Termglinchan V, Wu JC. Human Induced Pluripotent Stem Cell Models of Inherited Cardiomyopathies. Curr Opin Cardiol. 2014;29(3):214. doi:10.1097/HCO.0000000000000049
8. Matsa E, Burridge PW, Wu JC. Human Stem Cells for Modeling Heart Disease and for Drug Discovery. Sci Transl Med. 2014;6(239):239ps6. doi:10.1126/SCITRANSLMED.3008921
9. Lam CK, Wu JC. Disease modelling and drug discovery for hypertrophic cardiomyopathy using pluripotent stem cells: how far have we come? Eur Heart J. 2018;39(43):3893-3895. doi:10.1093/EURHEARTJ/EHY388
10. Matsa E, Burridge PW, Yu KH, et al. Transcriptome Profiling of Patient-Specific Human iPSC-Cardiomyocytes Predicts Individual Drug Safety and Efficacy Responses In Vitro. Cell Stem Cell. 2016;19(3):311-325. doi:10.1016/J.STEM.2016.07.006
11. Lan F, Lee AS, Liang P, et al. Abnormal Calcium Handling Properties Underlie Familial Hypertrophic Cardiomyopathy Pathology in Patient-Specific Induced Pluripotent Stem Cells. Cell Stem Cell. 2013;12(1):101-113. doi:10.1016/J.STEM.2012.10.010
12. Benzoni P, Campostrini G, Landi S, et al. Human iPSC modelling of a familial form of atrial fibrillation reveals a gain of function of If and ICaL in patient-derived cardiomyocytes. Cardiovasc Res. 2020;116(6):1147. doi:10.1093/CVR/CVZ217
13. Riley G, Syeda F, Kirchhof P, Fabritz L. An Introduction to Murine Models of Atrial Fibrillation. Front Physiol. 2012;3. doi:10.3389/FPHYS.2012.00296
14. Vincent F, Loria P, Pregel M, et al. Developing predictive assays: the phenotypic screening “rule of 3.” Sci Transl Med. 2015;7(293). doi:10.1126/SCITRANSLMED.AAB1201
15. Lee J, Termglinchan V, Diecke S, et al. Activation of PDGF pathway links LMNA mutation to dilated cardiomyopathy. Nat 2019 5727769. 2019;572(7769):335-340. doi:10.1038/s41586-019-1406-x
16. Brandao KO, Tabel VA, Atsma DE, Mummery CL, Davis RP. Human pluripotent stem cell models of cardiac disease: from mechanisms to therapies. Dis Model Mech. 2017;10(9):1039-1059. doi:10.1242/DMM.030320
17. Sala L, Yu Z, Oostwaard DW, et al. A new hERG allosteric modulator rescues genetic and drug‐induced long‐QT syndrome phenotypes in cardiomyocytes from isogenic pairs of patient induced pluripotent stem cells. EMBO Mol Med. 2016;8(9):1065. doi:10.15252/EMMM.201606260
18. Drawnel FM, Boccardo S, Prummer M, et al. Disease Modeling and Phenotypic Drug Screening for Diabetic Cardiomyopathy using Human Induced Pluripotent Stem Cells. Cell Rep. 2014;9(3):810-820. doi:10.1016/J.CELREP.2014.09.055
19. Blinova K, Dang Q, Millard D, et al. International Multisite Study of Human-Induced Pluripotent Stem Cell-Derived Cardiomyocytes for Drug Proarrhythmic Potential Assessment. Cell Rep. 2018;24(13):3582. doi:10.1016/J.CELREP.2018.08.079
20. Savoji H, Mohammadi MH, Rafatian N, et al. Cardiovascular disease models: A game changing paradigm in drug discovery and screening. Biomaterials. 2019;198:3-26. doi:10.1016/J.BIOMATERIALS.2018.09.036
21. Mills RJ, Parker BL, Quaife-Ryan GA, et al. Drug Screening in Human PSC-Cardiac Organoids Identifies Pro-proliferative Compounds Acting via the Mevalonate Pathway. Cell Stem Cell. 2019;24(6):895-907.e6. doi:10.1016/J.STEM.2019.03.009/ATTACHMENT/774AA312-FC80-4839-B0CE-2AD3F40B82F4/MMC4.XLSX
22. Giacomelli E, Meraviglia V, Campostrini G, et al. Human-iPSC-Derived Cardiac Stromal Cells Enhance Maturation in 3D Cardiac Microtissues and Reveal Non-cardiomyocyte Contributions to Heart Disease. Cell Stem Cell. 2020;26(6):862-879.e11. doi:10.1016/J.STEM.2020.05.004
23. Kathiresan S, Srivastava D. Genetics of Human Cardiovascular Disease. Cell. 2012;148(6):1242. doi:10.1016/J.CELL.2012.03.001
24. Matsa E, Rajamohan D, Dick E, et al. Fast Track: Editor’s Choice: Drug evaluation in cardiomyocytes derived from human induced pluripotent stem cells carrying a long QT syndrome type 2 mutation. Eur Heart J. 2011;32(8):952. doi:10.1093/EURHEARTJ/EHR073
25. Wu JC, Garg P, Yoshida Y, et al. Towards Precision Medicine with Human iPSCs for Cardiac Channelopathies. Circ Res. Published online August 30, 2020:653-658. doi:10.1161/CIRCRESAHA.119.315209

















Tell Us What You Think!